May 06, 2013 Abstract. Gene transcripts and transcript variants must be cloned to characterize gene function and regulation. However, obtaining full-length cDNAs with accurate sequences from the 5′ end through to the 3′ end can be challenging. Moultrie m-series digital game cameras user manuals. SMARTer® RACE 5’/3’ Kit User Manual (052617) takarabio.com Takara Bio USA, Inc. Page 4 of 30 I. Introduction The SMARTer RACE 5’/3’ Kit provides a method for performing both 5’- and 3’-rapid amplification of cDNA.
Smart Race Cdna Amplification Kit User Manual

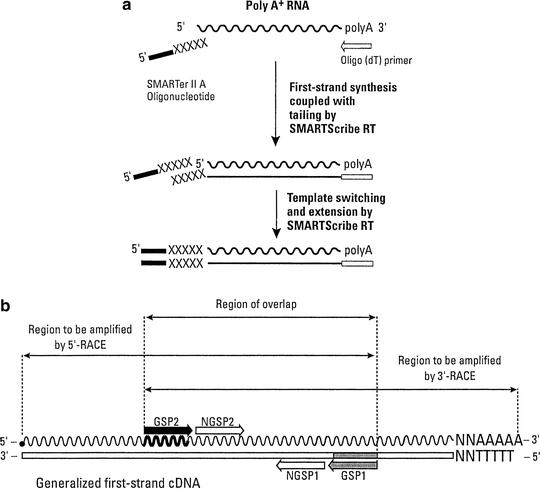
- SMART™ RACE cDNA Amplification Kit User Manual I. Introduction & Protocol Overview The SMART™ RACE cDNA Amplification Kit provides a method for performing both 5'- and 3'-rapid amplification of cDNA ends (RACE). This kit integrates our Marathon® cDNA Amplification Kit.
- What is the best RACE kit for amplify full cDNA genes? RACE kit (Rapid Amplification of cDNA ends) is used for amplification of 3' or 5' of cDNA. Attached with this email is the User.
- Instruction Manual 5´ RACE System for Rapid Amplification of cDNA Ends, Version 2.0 Catalog no. 18374-058 Version E 6 December 2004 50327.
Smarter Race Cdna Amplification Kit User Manual Free
- Gustincich S, Sandelin A, Plessy C, Katayama S, Simone R, Lazarevic D, Hayashizaki Y, Carninci P (2006) The complexity of the mammalian transcriptome. J Physiol 575:321–332PubMedCrossRefGoogle Scholar
- Hume DA (2008) Our evolving knowledge of the transcriptional landscape. Mamm Genome 19:663–666PubMedCrossRefGoogle Scholar
- Singer GA, Wu J, Yan P, Plass C, Huang TH, Davuluri RV (2008) Genome-wide analysis of alternative promoters of human genes using a custom promoter tiling array. BMC Genomics 9:349PubMedCrossRefGoogle Scholar
- Kim E, Goren A, Ast G (2008) Alternative splicing: current perspectives. Bioessays 30:38–47PubMedCrossRefGoogle Scholar
- Hiller M, Platzer M (2008) Widespread and subtle: alternative splicing at short-distance tandem sites. Trends Genet 24:246–255PubMedCrossRefGoogle Scholar
- Affymetrix/CSHL ENCODE Project (2009) Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457:1028–1032CrossRefGoogle Scholar
- Powell LM, Wallis SC, Pease RJ, Edwards YH, Knott TJ, Scott J (1987) A novel form of tissue-specific RNA processing produces apolipoprotein-B48 in intestine. Cell 50:831–840PubMedCrossRefGoogle Scholar
- Chen SH, Habib G, Yang CY, Gu ZW, Lee BR, Weng SA, Silberman SR, Cai SJ, Deslypere JP, Rosseneu M (1987) Apolipoprotein B-48 is the product of a messenger RNA with an organ-specific in-frame stop codon. Science 238:363–366PubMedCrossRefGoogle Scholar
- Carninci P (2006) Tagging mammalian transcription complexity. Trends Genet 22:501–510PubMedCrossRefGoogle Scholar
- Core LJ, Waterfall JJ, Lis JT (2008) Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322:1845–1848PubMedCrossRefGoogle Scholar
- Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M, Watts L, Booten SL, Graham M, McKay R, Subramaniam A, Propp S, Lollo BA, Freier S, Bennett CF, Bhanot S, Monia BP (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3:87–98PubMedCrossRefGoogle Scholar
- Carninci P (2009) Molecular biology: the long and short of RNAs. Nature 457:974–975PubMedCrossRefGoogle Scholar
- Hao S, Baltimore D (2009) The stability of mRNA influences the temporal order of the induction of genes encoding inflammatory molecules. Nat Immunol 10:281–288PubMedCrossRefGoogle Scholar
- Clontech (2009) SMARTer RACE cDNA amplification kit user manual. Clontech, Mountain View, CA, pp 1–33Google Scholar
- Kong W, Zhao JJ, He L, Cheng JQ (2009) Strategies for profiling microRNA expression. J Cell Physiol 218:22–25PubMedCrossRefGoogle Scholar
- Sdassi N, Silveri L, Laubier J, Tilly G, Costa J, Layani S, Vilotte JL, Le PF (2009) Identification and characterization of new miRNAs cloned from normal mouse mammary gland. BMC Genomics 10:149PubMedCrossRefGoogle Scholar
- Kodzius R, Kojima M, Nishiyori H, Nakamura M, Fukuda S, Tagami M, Sasaki D, Imamura K, Kai C, Harbers M, Hayashizaki Y, Carninci P (2006) CAGE: cap analysis of gene expression. Nat Methods 3:211–222PubMedCrossRefGoogle Scholar
- Olivarius S, Plessy C, Carninci P (2009) High-throughput verification of transcriptional starting sites by Deep-RACE. Biotechniques 46:130–132PubMedCrossRefGoogle Scholar
- Youngblood V, Taylor JV (2010) Sequencing PCR-amplified DNA in lipoprotein and cardiovascular disease research. In: Freeman LA (ed.) Methods in molecular biology: lipoproteins, 2nd edn, Walker JM (series ed.). Humana, Totowa, NJGoogle Scholar
- Diaw L, Youngblood V, Taylor JV (2010) Introduction to next-generation nucleic acid sequencing in cardiovascular disease research. In: Freeman LA (ed.) Methods in molecular biology: lipoproteins, 2nd edn, Walker JM (series ed.). Humana, Totowa, NJGoogle Scholar
- Wagner EM (2010) Monitoring gene expression: quantitative real-time RT-PCR. In: Freeman LA (ed.) Methods in molecular biology: lipoproteins, 2nd edn, Walker JM (series ed.). Humana, Totowa, NJGoogle Scholar
- Raghavachari N (2010) Microarray technology: basic methodology and application in clinical research for biomarker discovery in vascular diseases. In: Freeman LA (ed.) Methods in molecular biology: lipoproteins, 2nd edn, Walker JM (series ed.). Humana, Totowa, NJGoogle Scholar
- Koscielny G, Le TV, Gopalakrishnan C, Kumanduri V, Riethoven JJ, Nardone F, Stanley E, Fallsehr C, Hofmann O, Kull M, Harrington E, Boue S, Eyras E, Plass M, Lopez F, Ritchie W, Moucadel V, Ara T, Pospisil H, Herrmann A, Reich G, Guigo R, Bork P, Doeberitz MK, Vilo J, Hide W, Apweiler R, Thanaraj TA, Gautheret D (2009) ASTD: the alternative splicing and transcript diversity database. Genomics 93:213–220PubMedCrossRefGoogle Scholar
- Sabol SL, Brewer HB Jr, Santamarina-Fojo S (2005) The human ABCG1 gene: identification of LXR response elements that modulate expression in macrophages and liver. J Lipid Res 46:2151–2167PubMedCrossRefGoogle Scholar
- Shiraki T, Kondo S, Katayama S, Waki K, Kasukawa T, Kawaji H, Kodzius R, Watahiki A, Nakamura M, Arakawa T, Fukuda S, Sasaki D, Podhajska A, Harbers M, Kawai J, Carninci P, Hayashizaki Y (2003) Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc Natl Acad Sci U S A 100:15776–15781PubMedCrossRefGoogle Scholar
- Xu YX, Manley JL (2007) Pin1 modulates RNA polymerase II activity during the transcription cycle. Genes Dev 21:2950–2962PubMedCrossRefGoogle Scholar
